Richard Strasser
Molecular Plant Glycobiology
Orientation
Plant glycobiology, Plant cell biology, posttranslational modifications of proteins, intracellular transport and trafficking of proteins.
Molecular characterisation of the plant N-glycosylation pathway, characterisation of plant glycosyltransferases and glycosidases, biological function of N-glycans in plants, targeting and transport mechanisms of glycosylation enzymes, ER quality control systems, O-glycan engineering
Leader: Assoc. Prof. DI Dr. Richard Strasser
Projects
Selected Projects
- Functional characterization of complex N-glycans in plants
The main focus of our research is the investigation of the N-glycosylation pathway and the biological function of N-glycans in plants. During the last ten years we have characterized a set of mutants with clearly defined defects in N-glycan processing. These mutants are valuable tools to determine the role of individual glycoforms in a given process. We are especially interested to elucidate the biological significance of the final steps of N-glycan processing, which involve attachment of β1,2-linked xylose/core α1,3-linked fucose, removal of terminal GlcNAc residues by N-acetylhexosaminidases and the attachment of β1,3-galactose and α1,4-fucose to terminal GlcNAc residues.
- Investigation of glycan-dependent ER quality control systems in plants
In eukaryotes secreted proteins are subjected to a quality control system in the endoplasmic reticulum (ER), which retains newly synthesized polypeptides until they reach their correct conformation or targets potentially misfolded proteins for degradation. Both processes involve glycan-dependent chaperones and glycan trimming. While these quality control mechanisms are intensively studied in yeast and mammals, much less is known in plants. In this context we study the role of a family of A. thaliana class I α-mannosidases, which are involved in N-glycan trimming and ER-associated degradation of glycoproteins (ERAD).
- Investigation of mechanisms for subcellular localization of glycosylation enzymes
Plant N-glycan processing enzymes are arranged along the early secretory pathway, forming an assembly line in order to facilitate the step-by-step modification of oligosaccharides on glycoproteins. Thus, these enzymes provide excellent tools to study signals and mechanisms promoting their localization and retention in the ER and Golgi apparatus. Our main objective is the analysis of putative targeting motifs present in glycosidases and glycosyltransferases and the identification of interacting proteins to obtain new insights into this fundamental cellular process.
- O-glycan engineering in N. benthamiana
O-glycosylation is a common posttranslational modification of serine and threonine residues of secreted and membrane-bound proteins. In this project we extended our tool-box for glyco-engineering in the tobacco related species Nicotiana benthamiana towards the production of tailored mucin-type O-glycans on recombinant proteins.
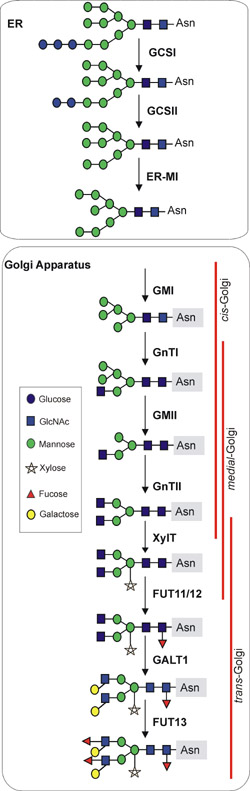
Figure 1. Processing steps of plant N-glycans in the ER and Golgi apparatus. GCSI: α-glucosidase I; GCSII: α-glucosidase II; ER-MI: ER-α-mannosidase I; GMI: Golgi-α-mannosidase I; GnTI: β1,2-N-acetylglucosaminyltransferase I; GMII: Golgi-α-mannosidase II; GnTII: β1,2-N-acetyl-glucosaminyltransferase II; XylT: β1,2-xylosyltransferase; FUT11/12: core α1,3-fucosyltransferase; GALT1: β1,3-galactosyltransferase; FUT13: α1,4-fucosyltransferase.
Publications
Authors' Self-Archiving Publications
- Dicker M, Strasser R. Using glyco-engineering to produce therapeutic proteins. Expert Opin Biol Ther. 2015;15(10):1501-1516. doi: 10.1517/14712598.2015.1069271. Epub 2015 Jul 14. PDF
- Strasser R. (2018). Protein Quality Control in the Endoplasmic Reticulum of Plants. Annu Rev Plant Biol 69: 147-172. doi: 10.1146/annurev-arplant-042817-040331
- Schoberer J, Shin YJ, Vavra U, Veit C, Strasser R. (2018). Analysis of Protein Glycosylation in the ER. Methods Mol Biol. 1691: 205-222. doi: 10.1007/978-1-4939-7389-7_16. PDF
- R. Strasser (2009) Localization of plant N‐glycan processing enzymes along the secretory pathway, Plant Biosystems - An International Journal Dealing with all Aspects of Plant Biology, 143:3, 636-642, DOI: 10.1080/11263500903233391. PDF
- Göritzer K, Strasser R. Glycosylation of Plant-Produced Immunoglobulins.Exp Suppl. 2021;112:519-543. doi: 10.1007/978-3-030-76912-3_16. PDF
- Margolin EA, Strasser R, Chapman R, Williamson AL, Rybicki EP, Meyers AE. Engineering the Plant Secretory Pathway for the Production of Next-Generation Pharmaceuticals. Trends Biotechnol. 2020 Sep;38(9):1034-1044. doi: 10.1016/j.tibtech.2020.03.004. PDF