Metabolomics of mycoparasitic interactions of Trichoderma atroviride and Botrytis cinerea
SUPERVISORS: Rainer SCHUHMACHER
PROJECT ASSIGNED TO: Kristina MISSBACH
Filamentous ascomycetes are prolific producers of secondary metabolites (SMs). Many natural bioactive products such as antibiotics, growth stimulants, flavors, and pigments are produced by microbes via secondary metabolism making fungal SMs an exciting topic, of not only scientific, but also commercial interest.
Trichoderma species are filamentous ascomycetes that are able to attack and parasitize on other living fungi. This survival strategy involves various mechanisms including the production of secondary metabolites which help Trichoderma to effectively compete against other (micro)organisms. While T. atroviride shows a great potential for the production of a diversity of SMs, most of the genes predicted to be associated with SM production however, are not expressed under routine laboratory conditions. One approach to induce the production of (novel) SMs is by mimicking the natural environment of the fungus, by co-cultivation with other microbes, assuming that cluster activation and subsequent metabolite production is mediated by chemical cues, released by the interaction partners.
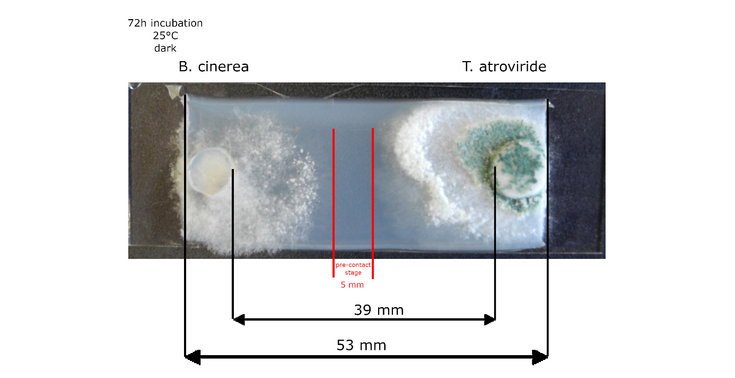

Picture: © Daniel Flatschacher, University of Innsbruck – interaction zone marked in red.
In this project we will combine confrontation assays between Trichoderma atroviride and Botrytis cinerea to provoke the formation of (novel) fungal secondary metabolites with stable isotope labeling (SIL) and LC-HRMS/MS. The whole secondary metabolomes of interacting T. atroviride and B. cinerea cultures will be investigated. The two fungi will be cultured in a miniaturized confrontation assay setup to maximize reproduce ability and sampling in the interaction zone. Fungal strains will be cultured under different labeling regimes using native (12C, 14N) or labeled (13C, 15N) nutrition sources. Labeling-specific isotope patterns, which will depend on the labeling strategy/purpose of the experiment will be automatically extracted from the raw data to assign all fungus derived metabolites by our in-house developed software MetExtract II. Single cultures and self-interactions (Botrytis-Botrytis, Trichoderma-Trichoderma) will serve as benchmark controls to identify interaction specific changes.
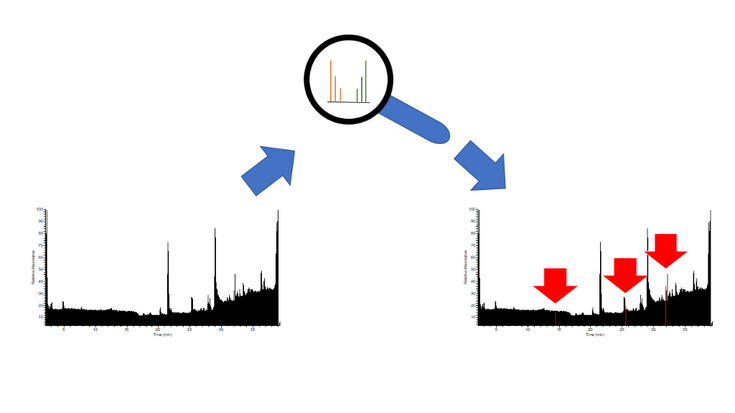
Picture 2: High resolution mass spectrometric (HRMS)-measurements result in an abundance of data. With stable isotope labeling, distinct isotope patterns can be recognized and are used to pick out relevant signals.
This work is part of a research project called “Chemical Cross-talk in mycoparasitic interactions (Chem-Talk)”, which is a cooperative research project of the BOKU university Vienna, the university of Innsbruck and the TU Wien.
