ATANASOVA Lab
Research Focus
Atanasova Lab is rooted in systems biology, exploring the complex, multi-layered networks that dictate how fungi interact with their environment and other organisms. Instead of studying individual genes in isolation, we adopt a holistic approach that integrates high-throughput data to map the "big picture" of fungal behavior.
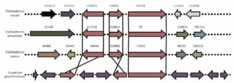
We focus on functional omics—specifically transcriptomics and proteomics—allowing us to analyze how entire sets of genes and proteins respond simultaneously during mycoparasitic interactions. By comparing these large-scale datasets across different fungal species and mutant strains, we can identify robustly regulated targets and the regulatory hierarchies that govern life-style switches. This systems-level perspective is essential for understanding lifestyle plasticity, as it reveals the coordinated signaling pathways, such as MAPK cascades, that act as central processing units for external stimuli.
Furthermore, our research links molecular data with phenotype profiling to bridge the gap between a fungus's genetic potential and its actual ecological performance. By investigating how bioactive metabolites and enzymatic activities function within the broader context of an ecosystem, we apply systems biology to understand how predictable biological control agents could work for sustainable agriculture. The integration of evolutionary analysis with functional genomics furthermore provides a comprehensive framework for understanding the intricate life strategies of filamentous fungi.
Atanasova’s research group explores the molecular biology and ecology of filamentous fungi, primarily focusing on how they interact with their environment and other organisms.
Our Lab integrates functional genomics, high-throughput phenotyping, and evolutionary analysis to understand how fungi adapt to different environments and host organisms.