Investigation and metabolic modelling of glycosylation in chinese hamster ovary (cho) cells
SUPERVISOR: Nicole BORTH
PROJECT ASSIGNED TO: Jerneja ŠTOR
Glycosylation, the attachment of sugar moieties, is a posttranslational modification which takes place in the endoplasmatic reticulum (ER) and the Golgi apparatus (GA). It does not follow a template but is the result of a complex network of enzymatic reaction (Figure 1) (Zhang et al., 2016). Proper glycosylation is a key quality parameter for biopharmaceuticals. The aim of this project is to develop analytical and computational methods that accurately predict the most important product quality parameter as function of the (bio-)process parameters.
The complexity of glycosylation is coupled with enzyme location in different compartments, enormous network of glycosylation enzymes, substrate preferences among the enzymes and redundancy among enzymatic activities. For this reason, it is difficult to investigate the process just by experimental work, and a suitable alternative approach is mathematical modeling. It complements experimental investigations in addressing the effects of cellular or process changes induced to increase recombinant protein productivity or maximize desired glycoform fractions within the glycoform population (Tejwani et al., 2018). A simplified outline of an already established glycosylation model from Sou et al., 2017, is presented with Figure 2.
In the course of the PhD thesis we will review the already existing glycosylation models and set the framework of new model. We will establish methods for the assessment of the glycosylation network in selected CHO cell line and carry out the experiments for the investigation of glycosylation. In this part we will focus on precise determination of glycosylation profile of the selected protein, precursor synthesis and expression of glycosylation enzymes. Established methods will allow precise determination of glycosylation network and rates of all steps of glycosylation in the chosen model cell lines. These data sets will then be used to validate the model and run simulations. A combined model that couples the existing GSMM with a kinetic model of protein production will improve predictions of productivity and will enable insights into the parameters that influence protein glycosylation by elucidating some of the underlying mechanisms.
The project is supported by the COMET center: acib -- Next Generation Bioproduction funded by BMK, BMDW, SFG, Standortagentur Tirol, Government of Lower Austria und Vienna Business Agency in the framework of COMET - Competence Centers for Excellent Technologies. The COMET-Funding Program is managed by the Austrian Research Promotion Agency FFG.
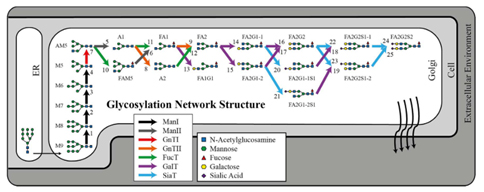
Figure 1: N-linked glycosylation network for immunoglobulin G in CHO-S. The corresponding stoichiometric model describes the flux balance among 25 glycosylation fluxes and 19 IgG glycoforms (excluding M9). The color of the arrows indicates the glycosylation enzyme that catalyzes the corresponding reaction (Hutter et al., 2017).
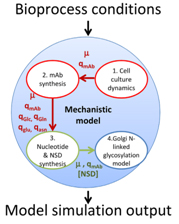
Figure 2: The modularity of the developed mathematical model and the interactions of each individual component within the overall model (Sou et al., 2017)
Hutter, S., Villiger, T. K., Brühlmann, D., Stettler, M., Broly, H., Soos, M., & Gunawan, R. (2017). Glycosylation flux analysis reveals dynamic changes of intracellular glycosylation flux distribution in Chinese hamster ovary fed-batch cultures. Metabolic Engineering, 43(June), 9–20. doi.org/10.1016/j.ymben.2017.07.005
Sou, S. N., Jedrzejewski, P. M., Lee, K., Sellick, C., Polizzi, K. M., & Kontoravdi, C. (2017). Model-based investigation of intracellular processes determining antibody Fc-glycosylation under mild hypothermia. Biotechnology and Bioengineering, 114(7), 1570–1582. doi.org/10.1002/bit.26225
Tejwani, V., Andersen, M. R., Nam, J. H., & Sharfstein, S. T. (2018). Glycoengineering in CHO Cells: Advances in Systems Biology. Biotechnology Journal, 13(3), 1–16. doi.org/10.1002/biot.201700234
Zhang, P., Woen, S., Wang, T., Liau, B., Zhao, S., Chen, C., Yang, Y., Song, Z., Wormald, M. R., Yu, C., & Rudd, P. M. (2016). Challenges of glycosylation analysis and control: An integrated approach to producing optimal and consistent therapeutic drugs. Drug Discovery Today, 21(5), 740–765. doi.org/10.1016/j.drudis.2016.01.006
